for each particle do // Independent
while particle is alive do // Dependent
Move particle to collision site
Process particle collision
This method of parallelism is very memory efficient, as the total number of particles that must be kept in memory at once is equivalent to the total number of active threads being run in the simulation. However, as there are many different types of collision events, the history-based model means that there is no natural SIMD style parallelism available for work happening between different threads.
An alternative simulation model is the "event-based" model. In this model, parallelism is instead expressed over different collision (or "event") types. To facilitate this, all particles in the simulation are stored in memory at once. Each event kernel is executed in parallel on vectors of particles that currently require that event to be executed:
Get vector of source particles
while any particles are alive do // Dependent
for each living particle do // Independent
Move particle to collision site
for each living particle do // Independent
Process particle collision
Sort/consolidate surviving particles
This method of parallelism requires more memory and requires an extra stream compaction kernel to sort and organize the particles periodically to ready them for the different event kernels. The benefit of this model is that kernels can potentially be execute in a SIMD manner and with higher cache efficiency due to the potential to sort particles by material and energy. On CPU architectures, the costs of sorting and buffering particles typically outweigh the benefits of the event-based model, but on accelerator architectures the tradeoff has been found to usually be more favorable.
XSBench represents the multipole macroscopic cross section lookup kernel. This kernel is responsible for adding together microscopic cross section data from all nuclides present in the material the neutron is travelling through, given a certain energy:
Macroscopic cross section data is typically required for multiple reaction channels "c", such as the total cross section, fission cross section, etc. This data is typically stored in point-wise data form for each nuclide. There are multiple ways of accessing this data in an efficient manner which will be discusses in this section.
This is the default "naive" method of performing macroscopic XS lookups. XS data is stored for a number of energy levels for each nuclide in the simulation problem. Different nuclides can have a different number of energy levels. For instance, U-238 usually has over 100k energy levels, whereas some other nuclides may only have a few thousand. The "Nuclide Grid" is composed of all nuclides in the problem, with a variable number of data points for each nuclide. Each data point is composed of the energy level and accompanying cross section data for multiple different reaction channels:
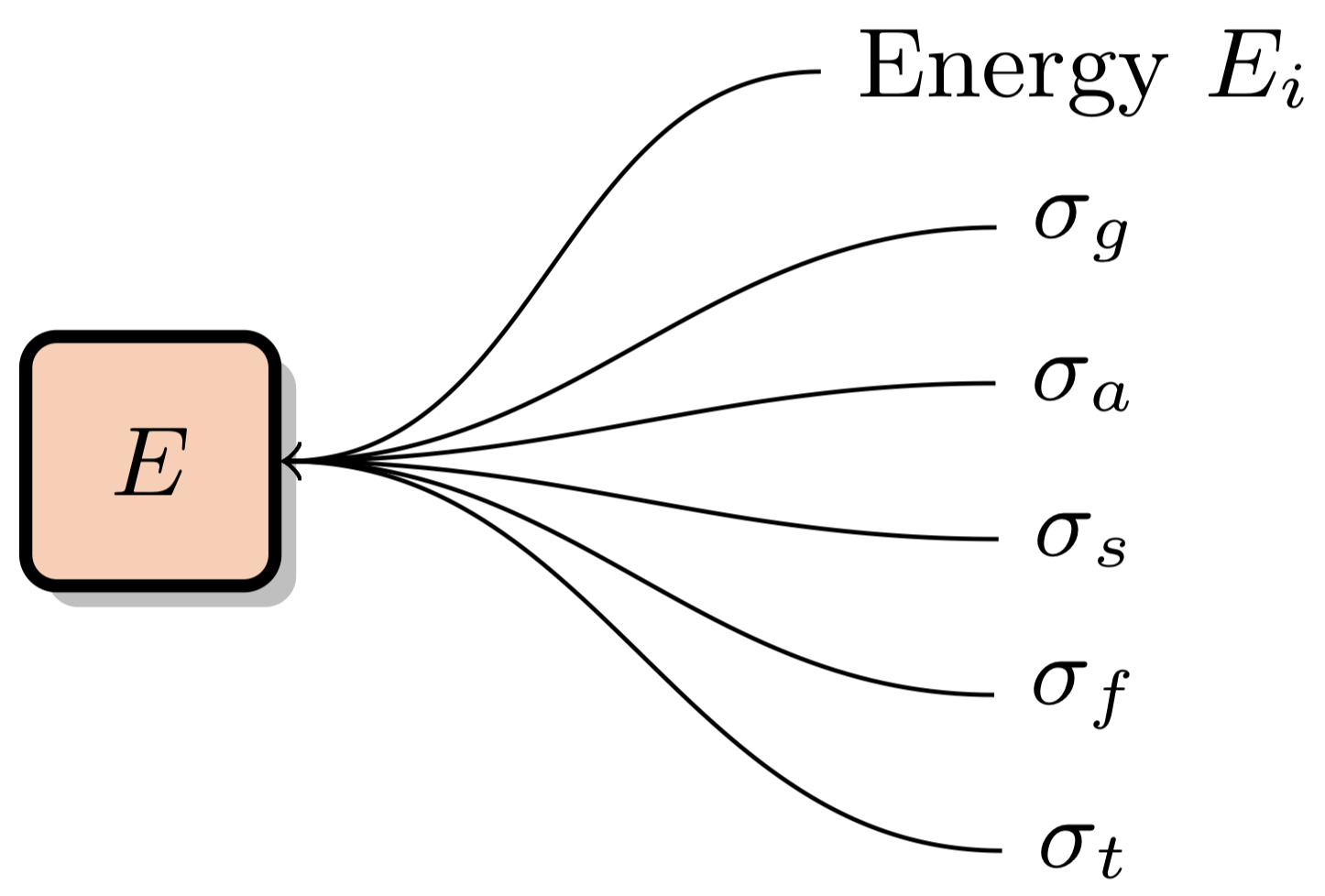
These XS data points are arranged into the nuclide grid:
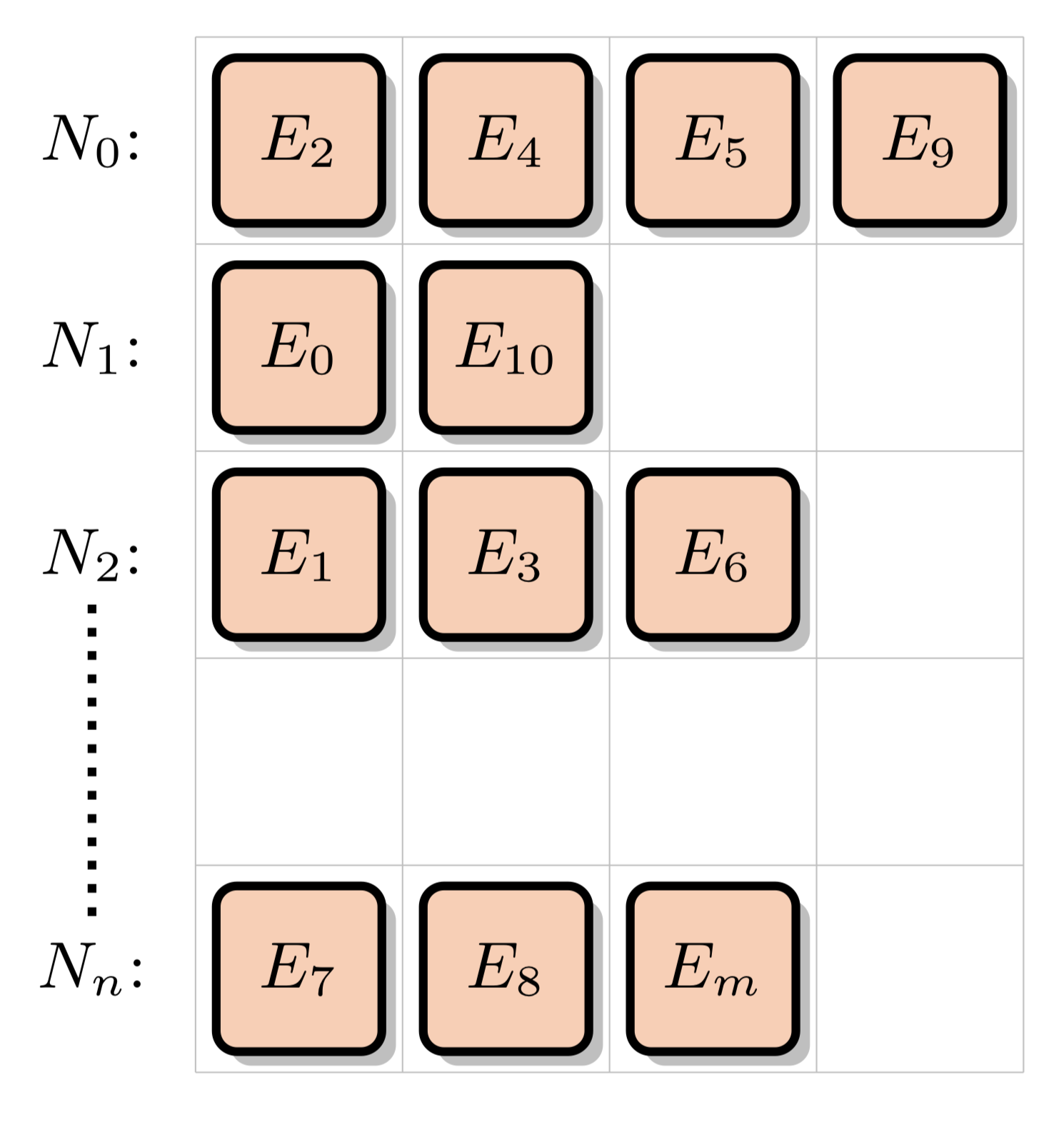
When assembling a macroscopic cross section data point, we will be accessing and interpolating data from the nuclide grid for a neutron travelling through a given material (composed of some number of nuclides) and at a given energy level. This will involve performing a binary search for each nuclide:
Nuclide_Grid_Search( Energy E, Material M ): macroscopic XS = 0 for each nuclide in M do: index = binary search to find E in nuclide grid interpolate data from grid[nuclide, index] macroscopic XS += data
This algorithm requires no extra memory usage beyond the minimum to represent the pointwise cross section data, but also requires that a binary search be run for each nuclide in the material the neutron is travelling through. In the case of depleted fuel, there can be 300+ nuclides, making this option computationally expensive.
One way of speeding up the nuclide grid search is to form a separate acceleration structure to reduce the number of binary searches that need to be performed. In the Unionized Energy Grid (EUG) method, a second grid is created with columns corresponding to the union of all energy levels from the nuclide grid. For each energy level (column) in the unionized grid, each row stores an index corresponding to the closest location in the nuclide grid for each nuclide corresponding that energy level:
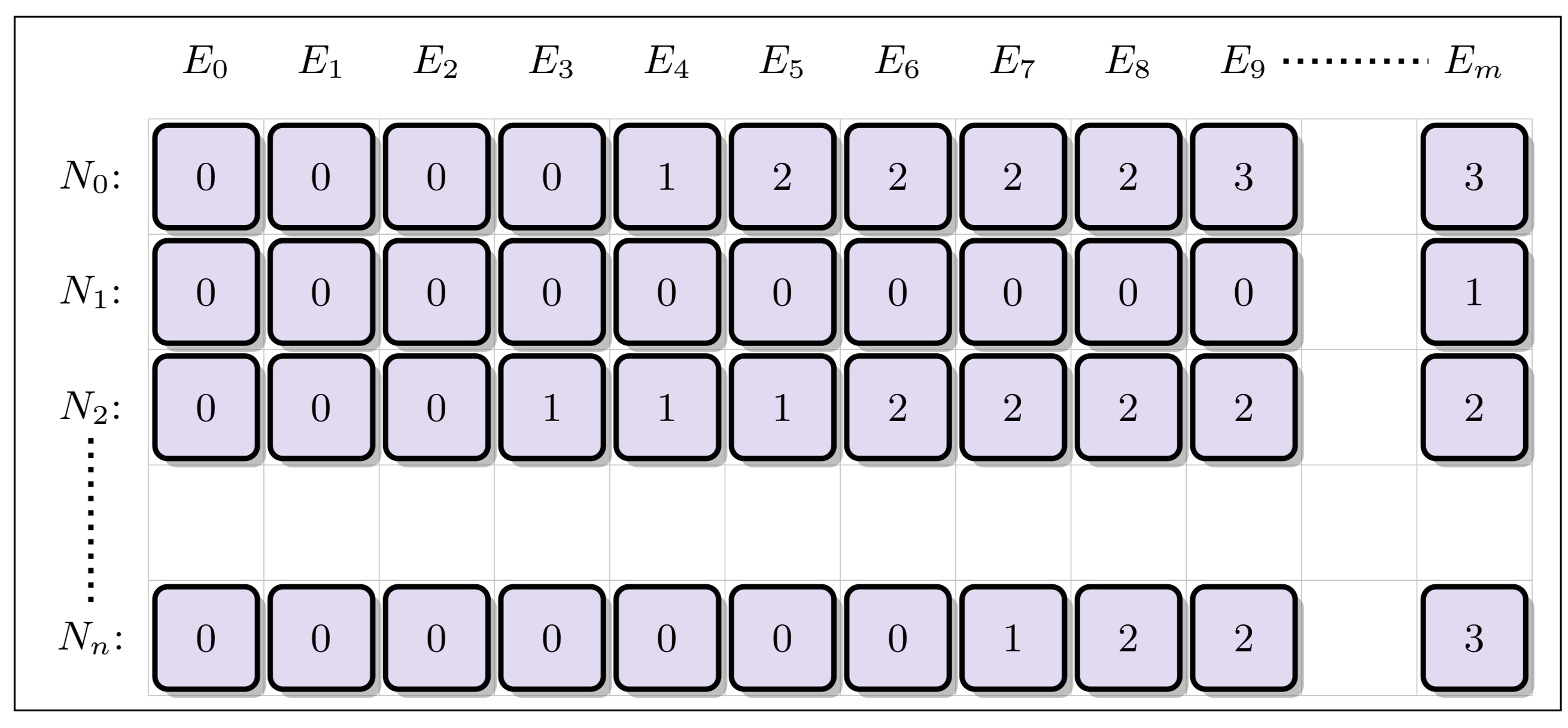
An alternative to the unionized energy grid is the logarithmic hash grid. This method takes in account the fact that while nuclides will be tabulated on grids containing different numbers of energy points, the points within each nuclide's grid will in general be spaced in (roughly) uniform manner in log space . Therefore, the nuclide grid is augmented with a separate acceleration structure similar to the unionized grid. However, the number of columns is capped at some number of bins spaced evenly in log space, with each row therefore corresponding to an approximate location within each nuclide's grid for that energy level. While the unionized grid points exactly to the correct index in the nuclide grid, the logarithmic hash grid points to only an approximate location below the true point -- meaning that a fast binary or iterative search must still be performed over the constrained area (typically only 10 or so elements in size):
Logarithmic_Hash_Grid_Search( Energy E, Material M ): macroscopic XS = 0 hash_index = grid_delta * (ln(E) - grid_minimum_energy) for each nuclide in M do: i_low = unionized_grid[nuclide, hash_index] i_high = unionized_grid[nuclide, hash_index+1] index = binary search in range(i_low, i_high) to find E in nuclide grid interpolate data from grid[nuclide, index] macroscopic XS += data
Compared to the unionized energy grid method, the logarithmic hash method uses far less memory but typically results in competitive performance. More details on this method can be found in the following publication:
Forrest B Brown. New hash-based energy lookup algorithm for monte carlo codes. Transactions of the American Nuclear Society, 111:659-662, 2014. permalink.lanl.gov/object/tr?what=info:lanl-repo/lareport/LA-UR-14-27037.
RSBench represents the multipole macroscopic cross section lookup kernel. This kernel is responsible for adding together microscopic cross section data from all nuclides present in the material the neutron is travelling through, given a certain energy:
Macroscopic cross section data is typically required for multiple reaction channels "c", such as the total cross section, fission cross section, etc.
Historically, cross section data has been stored in pointwise format, sometimes requiring in excess of 100,000 energy level data points be stored for a single nuclide. There are a variety of methods for performing cross section lookups on traditional pointwise data, as represented in the mini-app XSBench. However, a more memory and bandwidth efficient cross section representation method has recently been developed known as the "multipole" format that models the quantum mechanical resonances (or "poles") that underly the pointwise data. By this method, the resonances can be modeled mathematically and assembled on-the-fly while storing only a fraction of the data that is required for the traditional pointwise format. The tradeoff is that a greatly increased amount of floating point work must be performed when expanding the quantum mechanical "residues" into useable cross section data.
More information regarding the mathematics and equations used in the multipole
method can be found in:
C. Josey, P. Ducru, B. Forget, K. Smith, Windowed multipole for cross section
Doppler broadening, Journal of Computational Physics, Volume 307, 2016, Pages
715-727.
doi.org/10.1016/j.jcp.2015.08.013
Doppler broadening of resonance data requires evaluation of the complex error function, also known as the Faddeeva Functon. This function is not typically available as a language supplied or standard library intrinsic, so use of the multipole method requires implementing our own or adding a library dependency. Typical libraries that implement the Faddeeva function (e.g., the MIT Faddeeva Package written by Steven Johnson), break up the phase space of the function into many different areas, with each area using its own evaluation technique. This minimizes the number of floating point operations that must be performed, but can create a lot of branching which often precludes high SIMD efficiency. An alternative lightweight formulation is available, known as the Fast Nuclear Faddeeva (FNF) Function. FNF is a much simpler source implementation that only has one possible branch, allowing it to be specialized for use in complex phase space commonly seen in light water reactor simulations. FNF is therefore used in RSBench to minimize code complexity while providing acceptable accuracy for the use case of neutron transport.
If using the event-based model, we will be executing the lookup kernel in XSBench across all particles at once. While SIMD execution is possible using this method, typically issues can arrise that greatly reduce SIMD efficiency. In particular, different materials in the simulation have very different numbers of nuclides in them. For instance, spent fuel has 300+ nuclides, while moderator regions only have 10 or so nuclides. This creates a significant load imbalance across lanes in a SIMD vector, as some particles may only need a few iterations to complete all nuclides while others would need hundreds. Therefore, efficient SIMD execution of the event-based model is not possible without some optimizations.
One promising optimization for the event-based model is to perform a key-value sort of particles: first by material, and then by energy within each material. The first sort by material allows for adjacent particles in the vector to typically reside in the same type of material -- meaning that they will require the same number of nuclide lookup iterations. Then, the energy sort means that adjacent particles in the vector will be located close in energy space -- potentially allowing for many adjacent particles to access the same energy indices in each nuclide and therefore perform many or all of the same branching operations and read the same cache lines into memory at the same time. Once sorted, separate event kernels are then called for each material in the simulation. These two sorts can potentially boost both SIMD efficiency and cache efficiency, with effects being amplified as more particles are simulated at each event stage. The downside to this optimization is the introduction of the key-value particle sorting operations, which can be costly and potentially outweigh any gains due to improved SIMD efficiency and cache performance.
We have implemented this optimization in both the OpenMP threading and CUDA models. They are not enabled by default, but must be enabled with the "-k 1" or "-k 6" flags if running with OpenMP and CUDA respectively. These optimizations have not yet been implemented in the other programming models due to the lack of an efficient parallel sorting function being easily available without having to create an external library dependency. The CUDA version is disabled for SPEC CPU.
Copyright © 2026 Standard Performance Evaluation Corporation (SPEC®)